|
Viral Genomics & Ecoimmunology
VIRAL DISCOVERY. Leveraging biological collections and clinical sampling, I am using a combination of standard molecular approaches and targeted genomics to investigate the evolution of zoonotic viruses. The existence of diverse viral lineages in wildlife proximal to human habitation and agriculture highlights the critical need to expand investigations into viral diversity of natural reservoirs. With a focus on Afrotropical small mammals (bats, rodents, shrews), I am pairing viral data with ecological and immunological profiles to investigate these host-symbiont systems. I am also working with international collaborators to assess genomic diversity and epidemiological patterns of SARS-CoV-2 in West Africa. |
Press:
- "International collaboration strengthens global disease surveillance" (WARN-ID, 2023)
- "Andersen lab postdoc visits the National Public Health Institute of Liberia" (West African Research Network for Infectious Diseases, 2022)
- "By studying wild bats, alum seeks clues to COVID-19 immunity" (UChicago News, 2020)
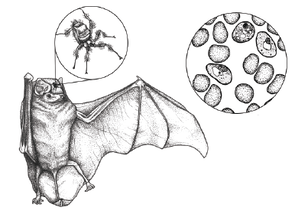 Figure by @MEMIllustration
Figure by @MEMIllustration
Animal Microbiomes
BATS & BIRDS. This collections-based, multiomics project is examining how adaptation to flight has shaped ecoimmunological conditions of the flighted vertebrate gut and subsequent evolution of associated microbiota. Initial sampling for this work has focused primarily on bats in the East African countries of Uganda and Kenya, transecting a broad range of ecosystems and habitat types, and also includes rodents, shrews, and birds from these regions. These samples are providing baseline data for a large number of species with diverse life histories, and allowi us to test hypotheses linking the bat microbiome, parasites, and pathogens.
Press
BATS & BIRDS. This collections-based, multiomics project is examining how adaptation to flight has shaped ecoimmunological conditions of the flighted vertebrate gut and subsequent evolution of associated microbiota. Initial sampling for this work has focused primarily on bats in the East African countries of Uganda and Kenya, transecting a broad range of ecosystems and habitat types, and also includes rodents, shrews, and birds from these regions. These samples are providing baseline data for a large number of species with diverse life histories, and allowi us to test hypotheses linking the bat microbiome, parasites, and pathogens.
Press
- "Blood-sucking flies may be following chemical cues..." (phys.org 2021)
- "The birds and the bats: Evolving to fly may have had big effect on gut microbiome" (SciGlow. 2020)
- "Gut Bacteria: How bats 'shift the paradigm'" (MedicalNewsToday, 2019)
- "Bat Microbiome is Unique Among Mammals" (Microbial Minutes, American Society for Microbiology, 2019)

CEPHALOPODS. In collaboration with the Marine Biological Laboratory (Woods Hole, MA), we are exploring the diversity of bacterial symbionts associated with cephalopods maintained in culture. Using 16S rRNA sequencing and fluorescence in situ hybridization, we identified a highly simplified microbiome in the European common cuttlefish, Sepia officinalis, comprising a single dominant Vibrio amplicon sequence variant (ASV) in the gastrointestinal tract of captive animals and a single dominant ASV in the family Piscirickettsia associated with the gills. We are now expanding this survey to include additional cephalopod species at MBL, as well as wild-caught individuals from the deep sea and are developing type cultures in the process with the goal of expanding the experimental toolkit for this highly promising system. Follow the Schmidt Ocean Institute for updates from our recent 2023 expeditions aboard the Falkor 'Too' and view the highlight reel from our first cruise with ROV SuBastian below,
Press:
- "Researchers found rare octopus nursery off the coast of Costa Rica" (npr.org, 2023)
- "Rare octopus nursery discovered nearly 2 miles below the ocean surface" (cnn.com, 2023)
- "Watch baby octopuses hatch from a surprising deep-sea nursery" (scientificamerican.org, 2023)
- "The cuttlefish may be flashy, but its microbiome is super simple" (sciencedaily.com, 2019)
|
DOLPHINS. In collaboration with the US Navy Marine Mammal Program, we are evaluating the microbial dynamics mediating the efficacy of fecal microbiome transplants for treatment of irritable bowel syndrome in common bottlenose dolphins (Tursiops truncatus). The underlying cause(s) of IBS in dolphins are not well known, and affected individuals may experience chronic symptoms that negatively impact quality of life and task performance. This multi-phase project is currently underway with completion anticipated in 2023. Read more: |
Ecology & Evolution of Malarial Parasites
Malaria parasites affect nearly all terrestrial vertebrates, and their impact on vertebrate hosts ranges from mild to severe, in some cases leading to extinction of entire species. Advances in molecular methods have revealed an astonishing degree of diversity among malaria parasites and closely-related haemosporidia, and host switching has occurred frequently throughout their evolutionary history. Without an accurate account of parasite diversity, we may only guess at the evolutionary history underlying current relationships. My research has focused primarily in the east African tropics, but has included Neotropical parasites as well. I use phylogenetic and comparative genomic approaches to resolve evolutionary relationships between malaria parasites of birds and mammals, and to explore the factors leading to diversification, host switching, and biogeographic distrbutions in haemosporidian parasites. Among the many African mammals I have tested, bats have proven to be consistent hosts of haemosporidian parasites - most of which represent new genetic lineages, and possibly new species. I continue to collect samples for haemosporidian analysis as I carry out my work on bats. View more work on Kenyan bats.
Press:
Malaria parasites affect nearly all terrestrial vertebrates, and their impact on vertebrate hosts ranges from mild to severe, in some cases leading to extinction of entire species. Advances in molecular methods have revealed an astonishing degree of diversity among malaria parasites and closely-related haemosporidia, and host switching has occurred frequently throughout their evolutionary history. Without an accurate account of parasite diversity, we may only guess at the evolutionary history underlying current relationships. My research has focused primarily in the east African tropics, but has included Neotropical parasites as well. I use phylogenetic and comparative genomic approaches to resolve evolutionary relationships between malaria parasites of birds and mammals, and to explore the factors leading to diversification, host switching, and biogeographic distrbutions in haemosporidian parasites. Among the many African mammals I have tested, bats have proven to be consistent hosts of haemosporidian parasites - most of which represent new genetic lineages, and possibly new species. I continue to collect samples for haemosporidian analysis as I carry out my work on bats. View more work on Kenyan bats.
Press:
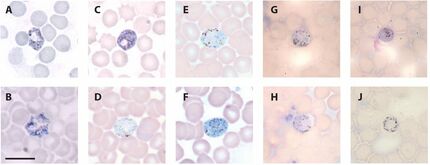
Micrographs of blood stage gametocytes of bat haemosporidians:
(A-C) Hepatocystis cf. epomophori macrogametocytes ex. Epomops franquetti; (D-E) Polychromophilus melanipherus microgametocytes and (F) macrogametocyte ex. Miniopterus natalensis; (G-H) Nycteria sp. microgametocyte, (I) macrogametocyte, and (J) gametocyte ex. Hipposideros cyclops.
LIFE HISTORY & PARASITISM. Prevalence and host-specificity of malaria parasites are highly variable across avian taxa. By comparing rates of parasitism across a broad range of bird species, we were able to assess the extent to which host life history traits and ecological factors are linked to infection by Plasmodium, Haemoproteus, or Leucocytozoon. To study the relationship between host traits and probability of haemosporidian infection, colleagues and I examined a broad range of bird species from 16 avian orders in Northern Malawi, discovering over 200 new parasite lineages in the process. This study was the first to test such associations in Afrotropical birds. This work is in collaboration with the Cornell Lab of Ornithology (Ithaca, NY) Field Museum of Natural History (Chicago, IL).
View this work in PLoS ONE.
View this work in PLoS ONE.
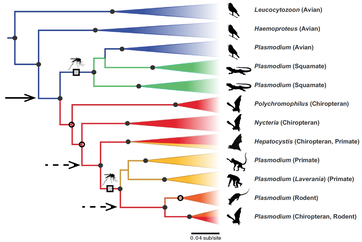
PHYLOGENETICS. To identify and describe malaria parasite diversity, I have sequenced the mitochondrial cytochrome b gene. While useful for differentiation between parasite lineages (see MalAvi database), this marker does not provide sufficient data for resolution of evolutionary relationships between malarial parasite lineages lineages of avian malarial parasites (Plasmodium, Haemoproteus, Leucocytozoon). To address this I collected additional data from nuclear and apicoplast genomes, allowing improved resolution for a diverse assemblage of malaria parasites I have identified in east African vertebrates.
View this work in Molecular Phylogenetics and Evolution.
View this work in Molecular Phylogenetics and Evolution.
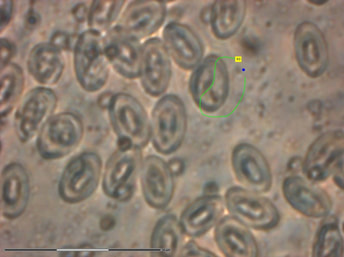
LASER CAPTURE & GENOMICS. Currently there are few publicly available genomes of Plasmodium parasites (see PlasmoDB), with no representation of avian Plasmodium species. My colleagues and I have used laser capture microdissection microscopy (LCMM) methods to obtain parasite isolates from birds for genomic sequencing and comparative analyses. Although this study recovered only a small fraction of genomic material, we were able to demonstrate that LCMM paired with whole genome amplification methods is a viable option for obtaining genomic material for comparative genomic analyses and the development of phylogenetic markers.
View this work in Parasitology Research.

